Keep getting this error when reading tomato leaves: "Exception Error, ReferenceError: ecsArray_flat_log is not defined"
Technical Support and Questions
Dhingra Lab
Feb 2019
Hello,
I am trying to get some readings from tomato plants in a greenhouse.
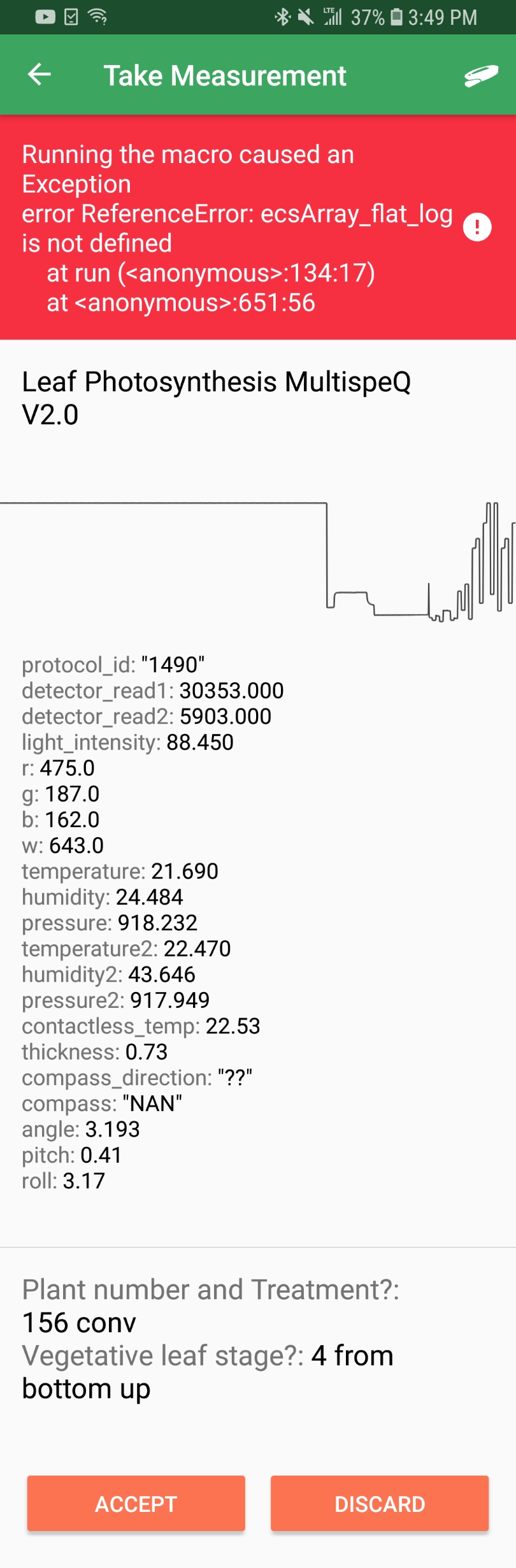
I keep receiving a red error message "Exception Error, ReferenceError: ecsArray_flat_log is not defined" (attached a screenshot of it). I searched for helping on the forums but I couldn't find any help.
Can anyone suggest some possible solutions?
Thank you!
David M. Kramer
Feb 2019
Hi! Is this Amit?
Can you post the firmware version and other settings? Go to console and type in "print_memory". Copy all text that appears and paste it here.
Dave
Dhingra Lab
Feb 2019
Hehe, no it's not Amit... we don't let him touch plants anymore. I'm one of his grad students. Thank you for the help Dr. Kramer!
Firmware: 2.0038
Memory dump:
{ "device_id":"32:21:3c:fb", "device_version":2, "compiled":["Sep 20 2018","09:30:55"], "mag_x_scale":-9999.00, "mag_y_scale":-9999.00, "mag_z_scale":-9999.00, "mag_x_offset":-9999.00, "mag_y_offset":-9999.00, "mag_z_offset":-9999.00, "light_slope_all":[0.614], "light_slope_r":[-0.583], "light_slope_g":[-0.551], "light_slope_b":[0.466], "light_yint":[-1.882], "thickness_a":16173.590, "thickness_b":-0.637, "thickness_c":6298.566, "mag_orientation":1.0, "open_position":34978.000, "closed_position":31924.000, "spad_scale":-9999.000, "blink_mode":4, "thickness_min":20869.400, "thickness_max":36290.602, "shutdown_time (s)":1800, "usb_on":1, "pix_pin":14, "par_to_dac_slope1": [8782.705,238855.516,13083.543,38828.016,10000.000,10000.000,74988.852,10000.000,10000.000,10000.000], "par_to_dac_slope2": [1223.51,77978.40,11179.30,15379.93,2000.00,2000.00,47558.15,2000.00,2000.00,2000.00], "par_to_dac_slope3": [-305.75,-147.63,-56.39,-313.22,-200.00,-200.00,-119.28,-200.00,-200.00,-500.00], "par_method":1, "par_tweek":-NAN } C38ACF17
Not sure why that date says Sept 20, 2018 because I've been trying to use the device these last couple days.
David M. Kramer
Feb 2019
Dhingra Lab: Hehe, no it's not Amit... we don't let him touch plants anymore. I'm one of his grad students. Thank you for the help Dr. Kramer! Firmware: 2.0038 Memory dump: { "device_id":"32:21:3c:fb", "device_version":2, "compiled":["Sep 20 2018","09:30:55"], "mag_x_scale":-9999.00, "mag_y_scale":-9999.00, "mag_z_scale":-9999.00, "mag_x_offset":-9999.00, "mag_y_offset":-9999.00, "...
Did you calibrate your magnetometer? It looks like you are getting NaN for compass. Try this: go to console. Type in "scan_i2c". When that is done, type in "calibrate_magnetometer" then slowly rotate the device in all directions making a sphere in the air.
Dhingra Lab
Feb 2019
I don't think I have calibrated the magnetometer. But I just tried and got error messages.
I tried to use those commands in the Console but just got {"error":"bad command"} each time.
David M. Kramer
Mar 2019
Hi Dhingra Lab. Let's get you running a more modern protocol. Can you try to run Photosynthesis RIDES?
Dhingra Lab
Apr 2019
That works Dr. Kramer! The RIDES protocol works much better without error.
I will run many more measurements in the future and I will keep you updated if there's any other issues.
Thank you for all the help!